16:9,将其用于汉坦病毒 => 画布:"科学社论信息图" 全局风格:"超写实3D,受冷冻电镜启发,纳米级微距布光,浅景深,病毒学美学" 标题:"VIRAL ARCHITECTURE: THE MULTISCALE LANDSCAPE OF [$VIRUS]" 布局引擎: 类型:"分层病毒形态学 + 组装路径" 连接器:"能量跃迁、对称轴、基因组–衣壳相互作用" 框架形状:"根据 [$VIRUS] 推断的二十面体/螺旋/复杂拓扑" 流向:"Genome → capsid → virion → host entry → replication cycle" 帧: f1_genome: 大小:"xs" 内容:"对 [$VIRUS] 基因组类型进行语义推断(RNA/DNA,ss/ds,分节段/非分节段)" UI标签:"Genome | Informational core | Nucleic acid topology" f2_capsid: 大小:"s" 内容:"对 [$VIRUS] 衣壳结构进行语义推断:二十面体、螺旋或复杂对称" UI标签:"Capsid | Symmetry | Structural proteins" f3_envelope: 大小:"m" 内容:"对 [$VIRUS] 包膜状态进行语义推断:脂质双层、糖蛋白、刺突、基质层" UI标签:"Envelope | Surface proteins | Host-derived membrane" f4_virion: 大小:"l" 内容:"对完整组装的 [$VIRUS] 病毒体进行语义推断:正确的形态、尺寸和表面拓扑" UI标签:"Virion | Infectious particle | Native morphology" f5_entry: 大小:"m" 内容:"对 [$VIRUS] 宿主进入机制进行语义推断:受体结合、融合、内吞或穿透" UI标签:"Entry | Receptor engagement | Membrane fusion or penetration" f6_replication: 大小:"m" 内容:"对 [$VIRUS] 复制策略进行语义推断:Baltimore分类逻辑、聚合酶使用、复制区室" UI标签:"Replication | Genome amplification | Viral factories" f7_assembly: 大小:"l" 内容:"对 [$VIRUS] 组装路径进行语义推断:衣壳形成、基因组包装、出芽或裂解" UI标签:"Assembly | Packaging | Maturation" f8_evolution: 大小:"xs" 内容:"对突变热点、准种群动力学和结构约束进行语义推断" UI标签:"Evolution | Mutational landscape | Fitness basins" 侧边面板: "衣壳三角剖分数(T)、螺旋节距、包膜糖蛋白密度、基因组长度、粒子直径、宿主范围图标" 光晕细节: "冷冻电镜剖面、表面静电、疏水性图谱、对称轴、受体结合足迹。 拒绝通用DNA双螺旋、药瓶或非病毒伪影。"
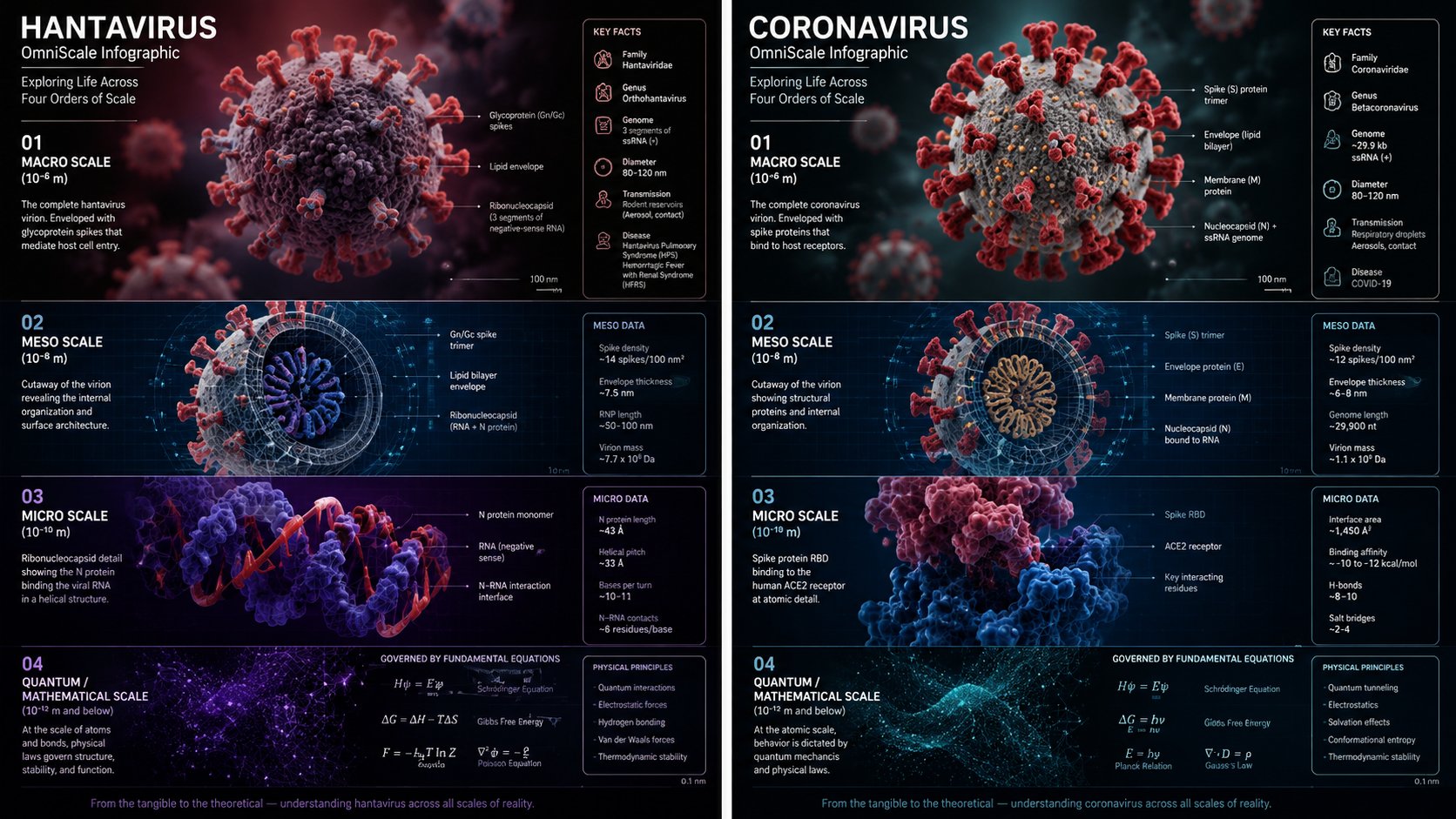
16:9, Do this for hantavirus => canvas: "Scientific Editorial Infographic" global_style: "Ultra-realistic 3D, cryo-EM inspired, nanoscopic macro-lighting, shallow DOF, virology aesthetic" title: "VIRAL ARCHITECTURE: THE MULTISCALE LANDSCAPE OF [$VIRUS]" layout_engine: type: "Hierarchical Viral Morphology + Assembly Pathway" connectors: "Energetic transitions, symmetry axes, genome–capsid interactions" frame_shape: "Icosahedral/Helical/Complex topology inferred from [$VIRUS]" flow_direction: "Genome → capsid → virion → host entry → replication cycle" frames: f1_genome: size: "xs" content: "Semantic inference of [$VIRUS] genome type (RNA/DNA, ss/ds, segmented/non-segmented)" ui_label: "Genome | Informational core | Nucleic acid topology" f2_capsid: size: "s" content: "Semantic inference of [$VIRUS] capsid architecture: icosahedral, helical, or complex symmetry" ui_label: "Capsid | Symmetry | Structural proteins" f3_envelope: size: "m" content: "Semantic inference of [$VIRUS] envelope state: lipid bilayer, glycoproteins, spikes, matrix layer" ui_label: "Envelope | Surface proteins | Host-derived membrane" f4_virion: size: "l" content: "Semantic inference of fully assembled [$VIRUS] virion: correct morphology, dimensions, and surface topology" ui_label: "Virion | Infectious particle | Native morphology" f5_entry: size: "m" content: "Semantic inference of [$VIRUS] host entry mechanism: receptor binding, fusion, endocytosis, or penetration" ui_label: "Entry | Receptor engagement | Membrane fusion or penetration" f6_replication: size: "m" content: "Semantic inference of [$VIRUS] replication strategy: Baltimore class logic, polymerase usage, replication compartments" ui_label: "Replication | Genome amplification | Viral factories" f7_assembly: size: "l" content: "Semantic inference of [$VIRUS] assembly pathway: capsid formation, genome packaging, budding or lysis" ui_label: "Assembly | Packaging | Maturation" f8_evolution: size: "xs" content: "Semantic inference of mutation hotspots, quasispecies dynamics, and structural constraints" ui_label: "Evolution | Mutational landscape | Fitness basins" side_panel: "Capsid triangulation numbers (T), helical pitch, envelope glycoprotein density, genome length, particle diameter, host range icons" halo_details: "Cryo-EM cross-sections, surface electrostatics, hydrophobicity maps, symmetry axes, receptor-binding footprints. Reject generic DNA helices, pill bottles, or non-viral artifacts."